Molecular Pathology
TEAM LEADER
Hugues DE THÉ
Professor
MEMBERS
ABDOLLAHZADEH Mahraz Master
mahraz.abdollahzadeh@etu.u-paris.fr
AYAKA Paolo Doctorant
paolo.ayaka@inserm.fr
DIRAMI Thassadite IR, UPC
thassadite.dirami@inserm.fr
ESNAULT Cécile CRCN, INSERM
cecile.esnault@inserm.fr
GEOFFROY Marie-Claude CRCN, CNRS
marie-claude.geoffroy@inserm.fr
KAWAKITA Michiko IR, UPC
michiko.kawakita@univ-paris-diderot.fr
MARMUSE Adèle Master
adele.marmuse@ens.psl.eu
RÉROLLE Domitille Doctorante
domitille.rerolle@college-de-france.fr
SOILIHI Hassane T, CNRS
hassane.soilihi@univ-paris-diderot.fr
TURAN Selim Bio-Informaticien
selim.turan@inserm.fr
WU Hsin-Chieh PostDoc
cjhsinchiehwu@gmail.com
ZHU Jun CRCN, INSERM
jun.zhu@univ-paris-diderot.fr
COLLÈGE DE FRANCE MEMBERS
ABOU-GHALI Majdouline PostDoc
majdouline.abou-ghali@college-de-france.fr
Pierre BERCIER ATER, Collège de France
pierre.bercier@college-de-france.fr
BERTHIER Caroline IE, Collège de France
caroline.berthier@college-de-france.fr
FERHI Omar IE, INSERM
omar.ferhi@college-de-france.fr
SITTLER Annie CRCN, CNRS
annie.sittler@college-de-france.fr
OUR TRUSTEES
OUR PARTNERS

APL pathogenesis
Acute Promyelocytic Leukaemia, APL, is a simple form of cancer driven by the PML/RARA fusion protein. PML/RARA is a transcriptional repressor that inhibits myeloid differentiation and enhances the survival and proliferation of early myeloid progenitors. PML/RARA also disrupts PML nuclear bodies, illustrating a now classic example of altered nuclear organisation and oncogenesis (de Thé, 2018; de Thé et al., 2017; Lallemand-Breitenbach and de Thé, 2018).
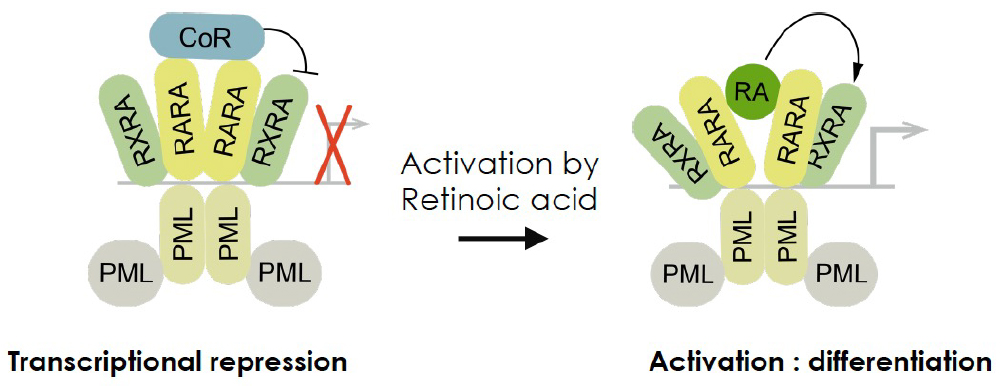
Two clinically highly efficient drugs, retinoic acid (RA) and arsenic were discovered in China. RA-induced reversion of PML-RARA transcriptional repression and gene re-activation was thought to be the molecular basis for the therapeutic effect of RA (Fig. 1). Yet, this model could not explain the action of arsenic (Zhu et al., 2002). Remarkably, we discovered that both RA and arsenic initiate PML/RARA degradation (Fig. 2). Although empirically discovered, these constitute oncogene-targeted therapies and APL has become the paradigm for efficient therapy through oncoprotein hyper-catabolism.
 RA-induced PML/RARA degradation and re-activation, yields in vivo differentiation, which is responsible for the massive tumor debulking. Yet, several studies, including our own, have shown that this is insufficient for prolonged remissions and cures (Nasr et al., 2008). In contrast, the combination of RA and arsenic can yield APL cure, in mouse models as well as in patients (Lallemand-Breitenbach et al., 1999; Lo-Coco et al., 2013). Mechanistically, APL eradication in mice models requires PML (Ablain et al., 2014) and accordingly, hotspot mutations in the normal PML protein were found in therapy-resistant patients (Lehmann-Che et al., 2014). It is presumed that restoration of PML nuclear bodies drives senescence of remaining leukemia-initiating APL cells. We are thus performing structure/function analyses of PML in the context of APL therapy.
RA-induced PML/RARA degradation and re-activation, yields in vivo differentiation, which is responsible for the massive tumor debulking. Yet, several studies, including our own, have shown that this is insufficient for prolonged remissions and cures (Nasr et al., 2008). In contrast, the combination of RA and arsenic can yield APL cure, in mouse models as well as in patients (Lallemand-Breitenbach et al., 1999; Lo-Coco et al., 2013). Mechanistically, APL eradication in mice models requires PML (Ablain et al., 2014) and accordingly, hotspot mutations in the normal PML protein were found in therapy-resistant patients (Lehmann-Che et al., 2014). It is presumed that restoration of PML nuclear bodies drives senescence of remaining leukemia-initiating APL cells. We are thus performing structure/function analyses of PML in the context of APL therapy.
PML is the key organiser of PML nuclear bodies (NBs), nuclear matrix-associated small spherical compartments, which recruit an ever-growing number of partner proteins (Lallemand-Breitenbach and de Thé, 2018) (Fig. 3). PML and NBs fine-tune a variety of processes (senescence, metabolism, self-renewal, apoptosis), most likely through facilitation of partner protein post-translational modifications, resulting in partner sequestration, activation or degradation. PML NBs are directly targeted by arsenic which enforces NB biogenesis through direct binding (Jeanne et al., 2010; Lallemand-Breitenbach et al., 2001; Zhu et al., 1997). We have recently discovered the structural basis for Arsenic binding onto a specific site of PML, addressing a long-standing issue in the field (Bercier et al., 2023). PML NBs are regulated by multiple cellular stresses: viral infection, DNA-damage, transformation, oxidative stress. Conversely, redox signalling and response to oxidative stress are profoundly altered in Pml-/- mice (Niwa-Kawakita et al., 2017).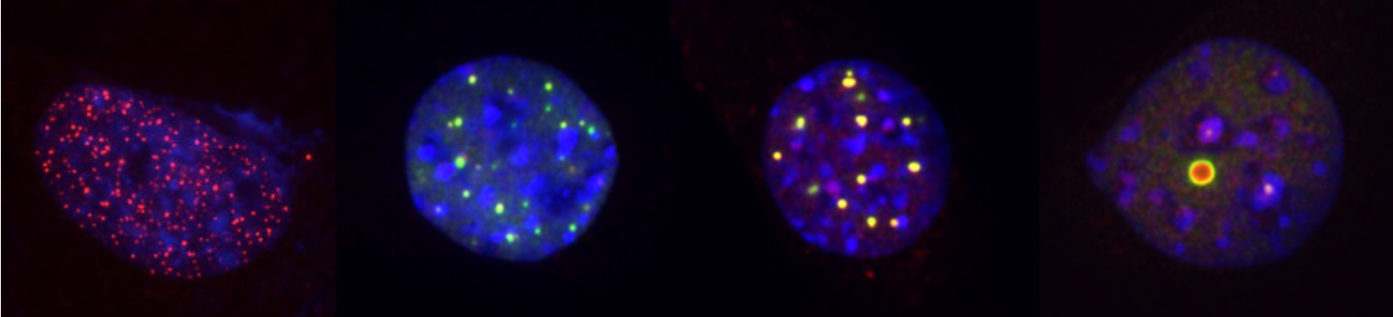
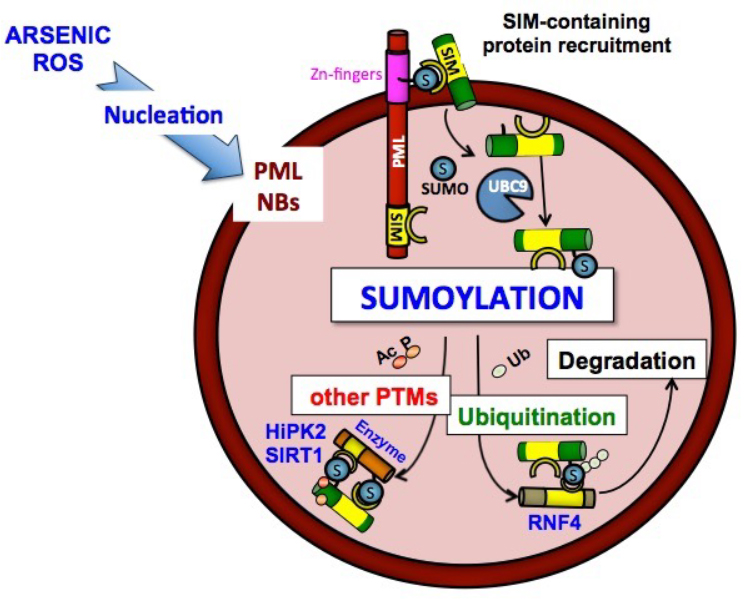
PML was the first protein identified as a major target of conjugation by the SUMO ubiquitin-like peptide. PML sumoylation drives recruitment of partners, through their SUMO-interacting motif (SIM) (Sahin et al., 2014). We have formally demonstrated that PML nuclear body promote sumoylation by concentrating UBC9 and its substrates (Tessier et al., 2022). Remarkably, in mESC, the phenotypes of Pml and Ubc9 knockouts are strikingly similar, implying that, at least in this cell-type, sumoylation efficiency relies on Pml presence. Some PML splice variants bind ARF, a UBC9 cofactor (Ivanschitz et al., 2015). Thus, PML NBs likely serve as catalytic chambers for sumoylation that recruit the enzyme (UBC9), the substrates (SUMOs and partner proteins) and cofactors (ARF) to ensure efficient, stress-inducible, SUMO conjugation and protein degradation. Indeed, poly- or multi-sumoylation can drive recruitment of proteins, such as RNF4, which couple recognition of SUMO (through their SIMs) to poly-ubiquitination. Similar to PML, some partner proteins are subsequently degraded in PML-dependent manner. Work from multiple labs have shown that this SUMO-initiated, ubiquitin-enforced degradation pathway is important, in particular for stress adaptation.
 Taken the key roles of PML in APL therapy response, we are interested in discovering other conditions where PML may similarly drive therapy response. One of them, myeloproliferative neoplasms with Jak2 mutation, was recently identified (Dagher et al., 2021). In NPM1c-associated AML, we have also demonstrated that the oncogene disrupts PML NBs and that Actinomycin D therapy restores them, identifying a situation with striking similarities to APL (Wu et al., 2021). PML NBs likely constitute a common effector pathway in therapy response of many leukemia (Rerolle and de Thé, 2023). Others diseases, including AML response to standard chemotherapy, are currently under study, as part of an ERC-funded project, PML-THERAPY.
Taken the key roles of PML in APL therapy response, we are interested in discovering other conditions where PML may similarly drive therapy response. One of them, myeloproliferative neoplasms with Jak2 mutation, was recently identified (Dagher et al., 2021). In NPM1c-associated AML, we have also demonstrated that the oncogene disrupts PML NBs and that Actinomycin D therapy restores them, identifying a situation with striking similarities to APL (Wu et al., 2021). PML NBs likely constitute a common effector pathway in therapy response of many leukemia (Rerolle and de Thé, 2023). Others diseases, including AML response to standard chemotherapy, are currently under study, as part of an ERC-funded project, PML-THERAPY.
We are now interested in identifying the mechanisms and pathways that ensure definitive APL clearance downstream of PML and their possible mechanistic relations to sumoylation. Current work on APL also involves exploration of the basis for APL initiation through deregulated retinoic acid signaling. Indeed, the rare APL variants not expressing PML/RARA constantly involve retinoic acid receptors (Geoffroy and de Thé, 2020). Interestingly, a sizable fraction (up to 30%) of non-APL AMLs over-express RARA and exhibit some RA-sensitivity, including in clinical trials. The molecular bases for these effects remain obscure, precluding the identification of clinically relevant biomarkers and identification of drug combinations with curative activities.
Altogether, APL is not only a clinical success story with almost all patients cured by targeted therapies, but also one of the too rare examples where cure is mechanistically understood in molecular and cellular details.
Some key references
►Ablain, J., Rice, K., Soilihi, H., de Reynies, A., Minucci, S., and de Thé, H. (2014). Activation of a promyelocytic leukemia-tumor protein 53 axis underlies acute promyelocytic leukemia cure. Nature medicine 20, 167-174. doi: 10.1038/nm.3441.
►Bercier, P., Wang, Q.Q., Zang, N., Zhang, J., Yang, C., Maimaitiyiming, Y., Abou-Ghali, M., Berthier, C., Wu, C., Niwa-Kawakita, M., et al. (2023). Structural basis of PML/RARA oncoprotein targeting by arsenic unravels a cysteine rheostat controlling PML body assembly and function. Cancer Discov. 10.1158/2159-8290.CD-23-0453.
►Dagher, T., Maslah, N., Edmond, V., Cassinat, B., Vainchenker, W., Giraudier, S., Pasquier, F., Verger, E., Niwa-Kawakita, M., Lallemand-Breitenbach, V., et al. (2021). JAK2V617F myeloproliferative neoplasm eradication by a novel interferon/arsenic therapy involves PML. J Exp Med 218. 10.1084/jem.20201268.
►De Thé, H. (2018). Differentiation therapy revisited. Nat Rev Cancer 18, 117-127. 10.1038/nrc.2017.103.
►De Thé, H., Pandolfi, P.P., and Chen, Z. (2017). Acute Promyelocytic Leukemia: A Paradigm for Oncoprotein-Targeted Cure. Cancer Cell 32, 552-560. 10.1016/j.ccell.2017.10.002.
► Geoffroy, M.C., and de Thé, H. (2020). Classic and Variants APLs, as Viewed from a Therapy Response. Cancers (Basel) 12. 10.3390/cancers12040967.
► Ivanschitz, L., Takahashi, Y., Jollivet, F., Ayrault, O., Le Bras, M., and de Thé, H. (2015). PML IV/ARF interaction enhances p53 SUMO-1 conjugation, activation, and senescence. Proc Natl Acad Sci U S A 112, 14278-14283. 10.1073/pnas.1507540112.
► Jeanne, M., Lallemand-Breitenbach, V., Ferhi, O., Koken, M., Le Bras, M., Duffort, S., Peres, L., Berthier, C., Soilihi, H., Raught, B., and de Thé, H. (2010). PML/RARA oxidation and arsenic binding initiate the antileukemia response of As2O3. Cancer Cell 18, 88-98. S1535-6108(10)00241-2 [pii] 10.1016/j.ccr.2010.06.003.
► Lallemand-Breitenbach, V., and de Thé, H. (2018). PML nuclear bodies: from architecture to function. Curr Opin Cell Biol 52, 154-161. 10.1016/j.ceb.2018.03.011.
► Lallemand-Breitenbach, V., Guillemin, M.-C., Janin, A., Daniel, M.-T., Degos, L., Kogan, S.C., Bishop, J.M., and de Thé, H. (1999). Retinoic acid and arsenic synergize to eradicate leukemic cells in a mouse model of acute promyelocytic leukemia. J. Exp. Med. 189, 1043-1052.
► Lallemand-Breitenbach, V., Zhu, J., Puvion, F., Koken, M., Honore, N., Doubeikovsky, A., Duprez, E., Pandolfi, P.P., Puvion, E., Freemont, P., and de Thé, H. (2001). Role of Promyelocytic Leukemia (PML) Sumolation in Nuclear Body Formation, 11S Proteasome Recruitment, and As(2)O(3)-induced PML or PML/Retinoic Acid Receptor alpha Degradation. J Exp Med 193, 1361-1372.
► Lehmann-Che, J., Bally, C., and de The, H. (2014). therapy resistance in APL. New Engl. J. Med. 371, 1171-1172.
► Lo-Coco, F., Avvisati, G., Vignetti, M., Thiede, C., Orlando, S.M., Iacobelli, S., Ferrara, F., Fazi, P., Cicconi, L., Di Bona, E., et al. (2013). Retinoic acid and arsenic trioxide for acute promyelocytic leukemia. The New England journal of medicine 369, 111-121. 10.1056/NEJMoa1300874.
► Nasr, R., Guillemin, M.C., Ferhi, O., Soilihi, H., Peres, L., Berthier, C., Rousselot, P., Robledo-Sarmiento, M., Lallemand-Breitenbach, V., Gourmel, B., et al. (2008). Eradication of acute promyelocytic leukemia-initiating cells through PML-RARA degradation. Nat Med 14, 1333-1342. nm.1891 [pii] 10.1038/nm.1891.
► Niwa-Kawakita, M., Ferhi, O., Soilihi, H., Le Bras, M., Lallemand-Breitenbach, V., and de Thé, H. (2017). PML is a ROS sensor activating p53 upon oxidative stress. J Exp Med 214, 3197-3206. 10.1084/jem.20160301.
► Rerolle, D., and de Thé, H. (2023). The PML hub: An emerging actor of leukemia therapies. J Exp Med 220. 10.1084/jem.20221213.
► Sahin, U., Ferhi, O., Jeanne, M., Benhenda, S., Berthier, C., Jollivet, F., Niwa-Kawakita, M., Faklaris, O., Setterblad, N., de Thé, H., and Lallemand-Breitenbach, V. (2014). Oxidative stress-induced assembly of PML nuclear bodies controls sumoylation of partner proteins. The Journal of cell biology 204, 931-945. 10.1083/jcb.201305148.
► Tessier, S., Ferhi, O., Geoffroy, M.C., Gonzalez-Prieto, R., Canat, A., Quentin, S., Pla, M., Niwa-Kawakita, M., Bercier, P., Rerolle, D., et al. (2022). Exploration of nuclear body-enhanced sumoylation reveals that PML represses 2-cell features of embryonic stem cells. Nat Commun 13, 5726. 10.1038/s41467-022-33147-6.
► Wu, H.C., Rerolle, D., Berthier, C., Hleihel, R., Sakamoto, T., Quentin, S., Benhenda, S., Morganti, C., Wu, C.C., Conte, L., et al. (2021). Actinomycin D targets NPM1c-primed mitochondria to restore PML- driven senescence in AML therapy. Cancer Discov In press.
► Zhu, J., Chen, Z., Lallemand-Breitenbach, V., and de Thé, H. (2002). How acute promyelocytic leukemia revived arsenic. Nature Reviews on Cancer 2, 705-713.
► Zhu, J., Koken, M.H.M., Quignon, F., Chelbi-Alix, M.K., Degos, L., Wang, Z.Y., Chen, Z., and de Thé, H. (1997). Arsenic-induced PML targeting onto nuclear bodies: implications for the treatment of acute promyelocytic leukemia. Proc. Natl. Acad. Sci. USA 94, 3978-3983.
The group’s primary interest is to explore the connections between genetic events driving cancer development and response to therapy, particularly unconventional ones.
In acute promyelocytic leukemia (APL), we have made key contributions to the understanding of the basis for leukemic transformation by the PML/RARA fusion protein and arsenic or retinoic acid therapy. Our studies have revealed that both agents bind PML/RARA and thus constitute targeted therapies. We demonstrated the key role of PML/RARA degradation by retinoic acid and arsenic in APL responses and the central role of PML in its cure in mice or patients.
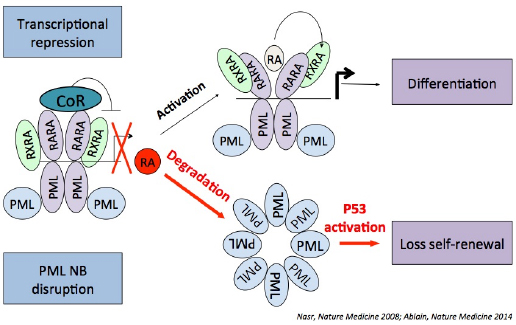

Over the years, the group has also invested very significant efforts to decipher the biochemical assembly and normal function(s) of PML NBs. Assembly of PML NB occurs through multiple sequential interactions within PML proteins, including ROS-regulated ones in the B2 box. Then SUMO conjugation of the PML shell allows recruitment of partner proteins, followed by their subsequent SUMO-conjugation. These proteins may be degraded by the SUMO-initiated, ubiquitin activated proteolysis pathway that we discovered by analyzing PML/RARA and PML degradation induced by arsenic. These sequential post-translational modifications contribute to stress adaptation.

Our current work, supported by the ERC (2019-2024), aims at harnessing PML nuclear bodies to activate a PML/p53 senescence axis in the context of other forms of leukemia treated with other drugs.
Collectively, our work is anchored at the rich interface between very basic issues of regulation of gene expression, protein post-translational modifications and cell biology, in the context of human cancer biology and novel therapies. Importantly, the strategies that derive from the models generated in the lab have been successfully implemented in APL patients.
Main publications – 2015-2021
► De Thé, H., P. P. Pandolfi and Z. Chen (2017). « Acute Promyelocytic Leukemia: A Paradigm for Oncoprotein-Targeted Cure. » Cancer Cell 32(5): 552-560.
► Niwa-Kawakita, M., O. Ferhi, H. Soilihi, M. Le Bras, V. Lallemand-Breitenbach* and H. de Thé* (2017). « PML is a ROS sensor activating p53 upon oxidative stress. » J Exp Med 214(11): 3197-3206.
► Ribet, D., V. Lallemand-Breitenbach, O. Ferhi, M. A. Nahori, H. Varet, H. de Thé* and P. Cossart* (2017). « Promyelocytic Leukemia Protein (PML) Controls Listeria monocytogenes Infection. » MBio 8(1).
► De Thé, H. (2018). « Differentiation therapy revisited. » Nat Rev Cancer 18(2): 117-127.
► Gachet, S., T. El-Chaar, D. Avran, E. Genesca, F. Catez, S. Quentin, M. Delord, G. Therizols, D. Briot, G. Meunier, L. Hernandez, M. Pla, W. K. Smits, J. G. Buijs-Gladdines, W. Van Loocke, G. Menschaert, I. Andre-Schmutz, T. Taghon, P. Van Vlierberghe, J. P. Meijerink, A. Baruchel, H. Dombret, E. Clappier, J. J. Diaz, C. Gazin, H. de Thé, F. Sigaux and J. Soulier (2018). « Deletion 6q Drives T-cell Leukemia Progression by Ribosome Modulation. » Cancer Discov 8(12): 1614-1631.
► Gaillard, C., S. Surianarayanan, T. Bentley, M. R. Warr, B. Fitch, H. Geng, E. Passegue, H. de Thé* and S. C. Kogan* (2018). « Identification of IRF8 as a potent tumor suppressor in murine acute promyelocytic leukemia. » Blood Adv 2(19): 2462-2466.
► Lallemand-Breitenbach, V. and H. de Thé (2018). « PML nuclear bodies: from architecture to function. » Curr Opin Cell Biol 52: 154-161.
► Lehmann-Che, J., C. Bally, E. Letouze, C. Berthier, H. Yuan, F. Jollivet, L. Ades, B. Cassinat, P. Hirsch, A. Pigneux, M. J. Mozziconacci, S. Kogan, P. Fenaux and H. de Thé (2018). « Dual origin of relapses in retinoic-acid resistant acute promyelocytic leukemia. » Nat Commun 9(1): 2047.
► Wang, P., S. Benhenda, H. Wu, V. Lallemand-Breitenbach, T. Zhen, F. Jollivet, L. Peres, Y. Li, S. J. Chen, Z. Chen, H. de Thé* and G. Meng* (2018). « RING tetramerization is required for nuclear body biogenesis and PML sumoylation. » Nat Commun 9(1): 1277.
► Esnault, C., R. Rahme, K. L. Rice, C. Berthier, C. Gaillard, S. Quentin, A. L. Maubert, S. Kogan and H. de Thé (2019). « FLT3-ITD impedes retinoic acid, but not arsenic, responses in murine acute promyelocytic leukemias. » Blood 133(13):1495-1506
► Gentric, G., Y. Kieffer, V. Mieulet, O. Goundiam, C. Bonneau, F. Nemati, I. Hurbain, G. Raposo, T. Popova, M. H. Stern, V. Lallemand-Breitenbach, S. Muller, T. Caneque, R. Rodriguez, A. Vincent-Salomon, H. de Thé, R. Rossignol and F. Mechta-Grigoriou (2019). « PML-Regulated Mitochondrial Metabolism Enhances Chemosensitivity in Human Ovarian Cancers. » Cell Metab 29(1): 156-173 e110.
► McKenzie, M. D., M. Ghisi, E. P. Oxley, S. Ngo, L. Cimmino, C. Esnault, R. Liu, J. M. Salmon, C. C. Bell, N. Ahmed, M. Erlichster, M. T. Witkowski, G. J. Liu, M. Chopin, A. Dakic, E. Simankowicz, G. Pomilio, T. Vu, P. Krsmanovic, S. Su, L. Tian, T. M. Baldwin, D. A. Zalcenstein, L. DiRago, S. Wang, D. Metcalf, R. W. Johnstone, B. A. Croker, G. I. Lancaster, A. J. Murphy, S. H. Naik, S. L. Nutt, V. Pospisil, T. Schroeder, M. Wall, M. A. Dawson, A. H. Wei, H. de Thé, M. E. Ritchie, J. Zuber and R. A. Dickins (2019). « Interconversion between Tumorigenic and Differentiated States in Acute Myeloid Leukemia. » Cell Stem Cell 25(2): 258-272 e259.
► Yuan, H., S. Gao, H. Chen, X. Liu, J. Zhou, H. de Thé and J. Zhu (2019). « Primitive macrophages are dispensable for HSPC mobilization and definitive hematopoiesis. » Blood 134(9): 782-784.
► Dagher, T., N. Maslah, V. Edmond, B. Cassinat, W. Vainchenker, S. Giraudier, F. Pasquier, E. Verger, M. Niwa-Kawakita, V. Lallemand-Breitenbach, I. Plo, J. J. Kiladjian, J. L. Villeval and H. de Thé (2021). « JAK2V617F myeloproliferative neoplasm eradication by a novel interferon/arsenic therapy involves PML. » J Exp Med 218(2) e20201268.
► Geoffroy, M. C., C. Esnault and H. de Thé (2021). « Retinoids in hematology: a timely revival? » Blood 137(18): 2429-2437.
► Wu, H. C., D. Rerolle, C. Berthier, R. Hleihel, T. Sakamoto, S. Quentin, S. Benhenda, C. Morganti, C. Wu, L. Conte, S. Rimsky, M. Sebert, E. Clappier, S. Souquere, S. Gachet, J. Soulier, S. Durand, J. J. Trowbridge, P. Benit, P. Rustin, H. El Hajj, E. Raffoux, L. Ades, R. Itzykson, H. Dombret, P. Fenaux, O. Espeli, G. Kroemer, L. Brunetti, T. W. Mak, V. Lallemand-Breitenbach, A. Bazarbachi, B. Falini, K. Ito, M. P. Martelli and H. de Thé (2021). « Actinomycin D targets NPM1c-primed mitochondria to restore PML-driven senescence in AML therapy. » Cancer Discov 11: 3198-3213.
► Wu, H. C., D. Rerolle and H. de Thé (2021). « PML/RARA destabilization by hyperthermia: a new model for oncogenic fusion protein degradation? » Blood Cancer Discov 2: 300-301.
► Tessier, S., O. Ferhi, M. C. Geoffroy, R. Gonzalez-Prieto, A. Canat, S. Quentin, M. Pla, M. Niwa-Kawakita, P. Bercier, D. Rerolle, M. Tirard, P. Therizols, E. Fabre, A. C. O. Vertegaal, H. de Thé* and V. Lallemand-Breitenbach* (2022). « Exploration of nuclear body-enhanced sumoylation reveals that PML represses 2-cell features of embryonic stem cells. » Nat Commun 13(1): 5726.
► Wu, H. C. and H. de Thé (2022). « In APL, noncoding mutations and SNP converge on WT1. » Blood 140(10): 1060-1061.
► Sebert, M., S. Gachet, T. Leblanc, A. Rousseau, O. Bluteau, R. Kim, R. Ben Abdelali, F. Sicre de Fontbrune, L. Maillard, C. Fedronie, V. Murigneux, L. Bellenger, N. Naouar, S. Quentin, L. Hernandez, N. Vasquez, M. Da Costa, P. H. Prata, L. Larcher, M. de Tersant, M. Duchmann, A. Raimbault, F. Trimoreau, O. Fenneteau, W. Cuccuini, N. Gachard, N. Auger, G. Tueur, M. Blanluet, C. Gazin, M. Souyri, F. Langa Vives, A. Mendez-Bermudez, H. Lapillonne, E. Lengline, E. Raffoux, P. Fenaux, L. Ades, E. Forcade, C. Jubert, C. Domenech, M. Strullu, B. Bruno, N. Buchbinder, C. Thomas, A. Petit, G. Leverger, G. Michel, M. Cavazzana, E. Gluckman, Y. Bertrand, N. Boissel, A. Baruchel, J. H. Dalle, E. Clappier, E. Gilson, L. Deriano, S. Chevret, F. Sigaux, G. Socie, D. Stoppa-Lyonnet, H. de Thé, C. Antoniewski, D. Bluteau, R. Peffault de Latour and J. Soulier (2023). « Clonal hematopoiesis driven by chromosome 1q/MDM4 trisomy defines a canonical route toward leukemia in Fanconi anemia. » Cell Stem Cell 30(2): 153-170 e159
► Bercier, P., Q. Q. Wang, N. Zang, J. Zhang, C. Yang, Y. Maimaitiyiming, M. Abou-Ghali, C. Berthier, C. Wu, M. Niwa-Kawakita, T. Dirami, M. C. Geoffroy, O. Ferhi, S. Quentin, S. Benhenda, Y. Ogra, Z. Gueroui, C. Zhou, H. Naranmandura, H. de Thé* and V. Lallemand-Breitenbach* (2023). « Structural basis of PML/RARA oncoprotein targeting by arsenic unravels a cysteine rheostat controlling PML body assembly and function. » Cancer Discov (Online)
► Rerolle, D. and H. de Thé (2023). « The PML hub: An emerging actor of leukemia therapies. » J Exp Med 220(8).
Julien ABLAIN
PhD Student
Shirine BENHENDA
Postdoc
Juliane HALFTERMEYER
PhD Student
Lisa IVANSCHITZ
PhD Student
Florence JOLLIVET
IE
Morgane Le BRAS
MCU
Anne-Lise MAUBERT
IE
Muege OGRUNC
Postdoc
Laurent PERES
TCN, CNRS
Kim RICE
Postdoc
Umut SAHIN
Postdoc
Adeline VITALIANO-PRUNIER
Postdoc
Hao YUAN
Postdoc
Gaël FORTIN
Doctorant
Emmanuel LAPLANTINE
CR
Valérie LALLEMAND-BREITENBACH
DR
Fang QIU
Postdoc
Kassandra LANCHAIS
IE










