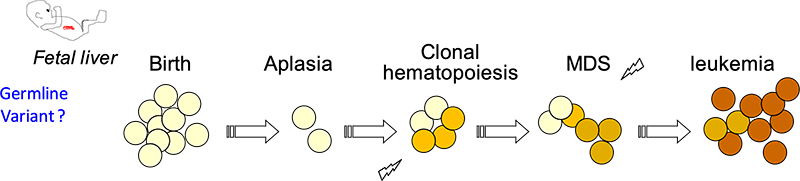
Our translational research aims to understand the drivers of clonal evolution, tumor progression and relapse in human acute leukemia. We especially study two human diseases prone to step-wise tumor progression, from preleukemia to leukemia, and from leukemia to relapse, Fanconi anemia (FA) and acute lymphoblastic leukemia (ALL).
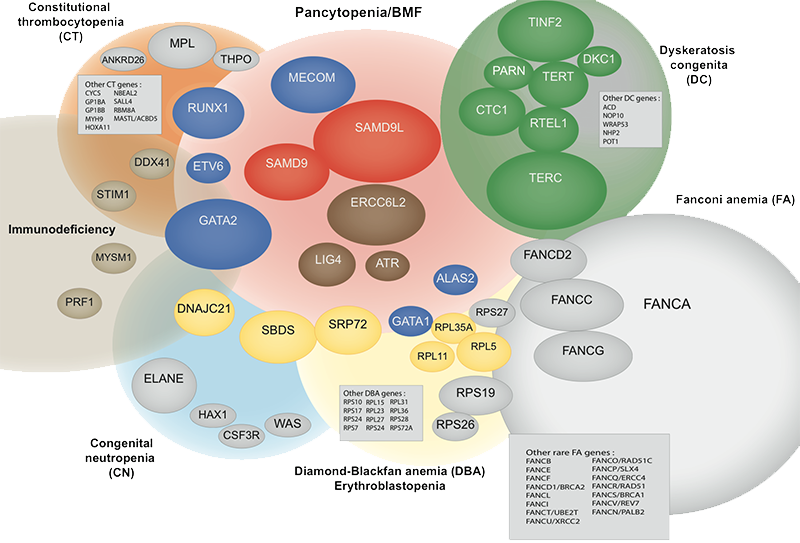
1/ Fanconi anemia (FA) and other inherited predispositions to bone marrow failure (BMF), myélodysplastic syndromes (MDS) and acute leukemia (AML)
Saint-Louis hospital is the national reference center for bone marrow failure diagnosis and treatment in France. Fanconi anemia (FA) is an inherited chromosomal instability disorder due to the inactivation of the FA/BRCA pathway. FA patients are predisposed to bone marrow failure and to develop MDS/leukemia during childhood or young adulthood. We have performed longitudinal genomic and functional analyses from primary samples of FA patients collected in the medical lab to identify the relevant mecanisms of BM progression. We aim to further model the disease in mice towards aplasia and MDS/AML. This research uncovered new insights in the pathogenesis of HSC attrition and subsequent oncogenesis in the context of constitutive genetic instability and aging. New BMF entities are being studied.
2/ Acute lymphoblastic leukemia, T or B-cell type
T and B-cell ALL (T-ALL) are heterogeneous hematopoietic malignancies derived from B or thymic progenitors. We used multiple genomic and expression analyses of T-ALL samples at diagnosis, at relapse and after engraftment in immunodeficient mice, to uncover the genetic landscape and functional pathways involved in tumor progression. In vitro and in vivo models are used to understand how the biological pathways are deregulated towards the driving forces that underlie tumor progression and relapse. These concepts and models are used for personalized stratification and therapy.
► Identification of new T-ALL oncogenic subtypes. Clonal evolution in T-ALL xenograft.
French referent lab for Fanconi anemia. Pathophysiology of hematopoietic failure and leukemogenesis in FA patients. New Bone marrow failure syndromes.
► Virus KSHV/HHV8 in the lymphoid proliferation Castleman’s disease.
► Sébert M, Passet M, Raimbault A, Rahmé R, Raffoux E, Sicre de Fontbrune F, Cerrano M, Quentin S, Vasquez N, Da Costa M, Boissel N, Dombret H, Peffault de Latour R, Socié G, Itzykson R, Fenaux P, Soulier J, Adès L, Clappier E. Germline DDX41 mutations define a significant entity within adult MDS/AML patients. Blood 2019 Oct 24;134(17):1441-1444.
► Nachmani D, Bothmer AH, Grisendi S, Mele A, Bothmer D, Lee JD, Monteleone E, Cheng K, Zhang Y, Bester AC, Guzzetti A, Mitchell CA, Mendez LM, Pozdnyakova O, Sportoletti P, Martelli MP, Vulliamy TJ, Safra M, Schwartz S, Luzzatto L, Bluteau O, Soulier J, Darnell RB, Falini B, Dokal I, Ito K, Clohessy JG, Pandolfi PP. Germline NPM1 mutations lead to altered rRNA 2’-O-methylation and cause dyskeratosis congenita. Nature Genetics 2019;51(10):1518-1529.
► Rio P, Navarro S, Wang W, Sánchez-Domínguez R, Pujol RM, Segovia J, Bogliolo M, Merino E, Wu N, Salgado R, Lamana ML, Yañez RM, Casado JA, Giménez Y, Román-Rodríguez FJ, Álvarez L, Alberquilla O, Raimbault A, Guenechea G, Lozano ML, Cerrato L, Hernando M, Gálvez E, Hladun R, Giralt I, Barquinero J, Galy A, García de Andoín N, López R, Catalá A, Schwartz JD, Surralles J, Soulier J, Schmidt M, Díaz de Heredia C, Sevilla J, Bueren JA. Successful Engraftment of Gene Corrected Hematopoietic Stem Cells in Non-conditioned Fanconi Anemia Patients. Nature Medicine 2019;25(9):1396-1401.
► Sarasin A, Quentin S, Droin N, Sahbatou M, Saada V, Auger N, Boursin Y, Dessen P, Raimbault A, Asnafi V, Schmutz JL, Taïeb A, Menck CFM, Rosselli F, Drieu La Rochelle L, Robert C, Sicre de Fontbrune F, Sébert M, Leblanc T, Kannouche P, De Botton S, Solary E, Soulier J. Familial predisposition to TP53/complex karyotype MDS and leukemia in DNA repair-deficient xeroderma pigmentosum. Blood 2019 Jun 20;133(25):2718-2724.
► Passet M, Boissel N, Sigaux F, Saillard C, Bargetzi M, Ba I, Thomas X, Graux C, Chalandon Y, Leguay T, Lengliné E, Konopacki J, Quentin S, Delabesse E, Lafage-Pochitaloff M, Pastoret C, Grardel N, Asnafi V, Lhéritier V, Soulier J, Dombret H, Clappier E; Group for Research on Adult ALL (GRAALL). PAX5 P80R mutation identifies a novel subtype of B-cell precursor acute lymphoblastic leukemia with favorable outcome. Blood 2019 Jan 17;133(3):280-284.
► Gachet S, El-Chaar T, Avran D, Genesca E, Catez F, Quentin S, Delord M, Thérizols G, Briot D, Meunier G, Hernandez L, Pla M, Smits WK, Buijs-Gladdines JG, Van Loocke W, Menschaert G, André-Schmutz I, Taghon T, Van Vlierberghe P, Meijerink JP, Baruchel A, Dombret H, Clappier E, Diaz JJ, Gazin C, de Thé H, Sigaux F, Soulier J. Deletion 6q drives T-cell leukemia progression by ribosome modulation. Cancer Discovery 2018 Dec;8(12):1614-1631.
► Bogliolo M, Bluteau D, Lespinasse J, Pujol R, Vasquez N, d’Enghien CD, Stoppa-Lyonnet D, Leblanc T, Soulier J*, Surrallés J*. Biallelic truncating FANCM mutations cause early-onset cancer but not Fanconi anemia. Genet Med 2018 Apr;20(4):458-463.
► Bluteau O, Sebert M, Leblanc T, Peffault de Latour R, Quentin S, Lainey E, Hernandez L, Dalle JH, Sicre de Fontbrune F, Lengline E, Itzykson R, Clappier E, Boissel N, Vasquez N, Da Costa M, Masliah-Planchon J, Cuccuini W, Raimbault A, De Jaegere L, Adès L, Fenaux P, Maury S, Schmitt C, Muller M, Blin N, Bruno B, Pellier I, Hunault M, Blanche S, Petit A, Leverger G, Michel G, Bertrand Y, Baruchel A, Socié G, Soulier J. A landscape of germline mutations in inherited bone marrow failure patients. Blood 2018 Feb 15;131(7):717-732.
► Domenech C, Maillard L, Rousseau A, Guidez F, Petit L, Pla M, Clay D, Guimiot F, Sanfilippo S, Jacques S, de la Grange P, Robil N, Soulier J, Souyri M. Studies in an early development window unveils a severe HSC defect in both murine and human Fanconi anemia. Stem Cell Reports 2018.
► Lesport E, Ferster A, Biver A, Roch B, Vasquez N, Jabado N, Vives FL, Revy P, Soulier J, de Villartay JP. Reduced recruitment of 53BP1 during interstrand crosslink repair is associated with genetically inherited attenuation of mitomycin C sensitivity in a family with Fanconi anemia. Oncotarget 2017;9(3):3779-3793.
► Bluteau D, Masliah-Planchon J, Clairmont C, Rousseau A, Ceccaldi R, Dubois d’Enghien C, Bluteau O, Cuccuini W, Gachet G, Peffault de Latour R, Leblanc T, Socié G, Baruchel A, Stoppa-Lyonnet D, D’Andrea AD, Soulier J. Biallelic inactivation of REV7 is associated with Fanconi anemia. J Clin Invest 2016;126(9):3580-4.
► Zhang H, Kozono DE, O’Connor KW, Vidal-Cardenas S, Rousseau A, Hamilton A, Moreau L, Gaudiano EF, Greenberger J, Bagby G, Soulier J, Grompe M, Parmar K, D’Andrea AD. TGF-β Inhibition Rescues Hematopoietic Stem Cell Defects and Bone Marrow Failure in Fanconi Anemia. Cell Stem Cell 2016;18(5):668-81.
► Peffault de Latour R, Soulier J. How we diagnose and t treat Fanconi anemia. Blood 2016;127(24):2971-9.