Identification and targeting of extrinsic regulators of myeloid malignancies
TEAM LEADER
Lina BENAJIBA, MD, PhD, HDR
CCA-INSERM-Bettencourt
CONTRIBUTORS
Lara ZAFRANI MD.PhD, HDR
Principal Investigator
Professor in Intensive Care Medicine
lara.zafrani@aphp.fr
Cécile CULEUX MSc
Lab manager
cecile.culeux@inserm.fr
Blandine ROUX PhD
Research Technician
blandine.roux@inserm.fr
Nina KACI MSc
Graduate student
nina.kaci@inserm.fr
Marta LUPERTO MD
Graduate Student
marta.luperto@inserm.fr
Federica NAMOR MSc
Graduate Student
federica.namor@inserm.fr
Hélène PASQUER MD
Graduate student
helene.pasquer@inserm.fr
Enora LE GOFF
Master Student
enora.le-goff@inserm.fr
Clarissa MUJACIC
Master student
c.mujacic@campus.unimib.it
OUR TRUSTEES
OUR PARTNERS

Myeloid neoplasms emerge from the accumulation of a number of somatic mutations within hematopoietic stem and progenitor cells. Presence of such mutations can lead to pre-leukemic states such as MPN (myeloproliferative neoplasms). Patients harboring MPN can transform into poor prognosis secondary acute myeloid leukemia (AML). Alternatively, AML can arise de novo without any pre-existing hematological disorder. AML is the most common acute leukemia diagnosed in adults and is accounted for approximately 25 000 new diagnosed cases per year in Europe. Despite the significant progress made in understanding AML pathogenesis over the last decades, our clinical progress in treating this disease has lagged behind resulting in a dismal 5-year overall survival (<25%). Extrinsic signals (microenvironment, treatment exposure and sepsis) play a key role in shaping clonal fitness within the bone marrow, therefore defining poorly understood myeloid oncogenesis paths. Applying cutting edge descriptive and functional large-scale technologies to physiopathologically relevant models, our team aims at better understanding these mechanisms in order to therapeutically leverage environment-driven clonal plasticity to prevent AML development and/or persistence.
BENAJIBA group
Accelerating therapeutic innovation to eradicate acute myeloid leukemia
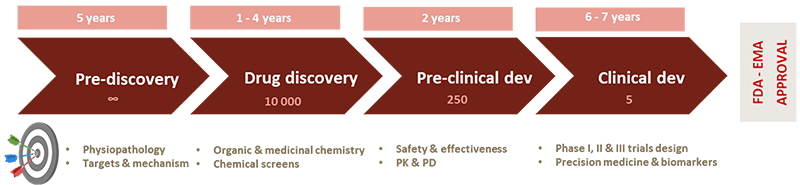
Developing new translational research strategies focused on the identification of druggable oncogenic targets is critical to pave the road for successful AML treatment. With an eye towards clinical translation, our research projects stem at the interface between biology, chemistry and clinical research in order to develop innovative methods, aiming to identify new therapeutic opportunities for patients with AML. To efficiently translate our target and drug discovery approaches to the clinic, our team works in close interaction with Saint-Louis hospital hematology clinic and early drug development department. We also seek to develop academic and industry interactions to accelerate the drug discovery journey in AML. Our discoveries may therefore contribute to novel and more efficacious treatments for this highly aggressive and lethal disease.
To accelerate therapeutic innovation in myeloid malignancies, our team focuses on three main axes of research:
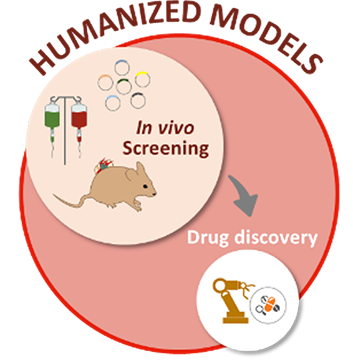
Developing physiologically relevant myeloid malignancies models
Relying on meaningful disease models is an essential prerequisite to identify therapeutic alternatives for patients with relapsed/refractory myeloid hematological malignancies. Driven in large part by therapeutic considerations, our team develops humanized myeloid malignancies mouse models, closely mimicking the physio-pathological conditions of malignant myeloid progenitors and leukemic cells growth within the bone marrow microenvironment. The engineering and optimization of such innovative and powerful mice models harboring ectopic patient derived neoplasms within humanized bone marrow niches will allow large scale therapeutic target discovery.
Unravelling the mechanistic underpinnings of MPN transformation into AML
Developing new translational approaches to identify key post-MPN AML development mechanisms is critical for hampering such dismal disease evolution. Leukemic transformation is a complex process involving intrinsic accumulation of genetic alterations in pre-leukemic cells, and cellular crosstalk between the hematopoietic cells and their niche. Therefore, clonal selection and subsequent leukemic transformation may result from the life-long therapeutic intervention needed to treat such chronic disorders, and/or from microenvironment-related signals.
To decipher the mechanism of pre-leukemic to leukemic state transition, we develop a comprehensive translational approach combining data-mining of an extensively annotated database, in vitro and in vivo models and large-scale functional and descriptive genomic approaches. We aim at highlighting potential therapy related selection of highly detrimental mutations and deciphering the mechanisms underlying such treatment-induced clonal selection. In parallel, we also develop powerful in vivo functional screening approaches within physiopathologically relevant MPN models, aiming to discover and validate novel niche-hematopoietic crosstalk transformation promoting pathways.
Our discoveries will thus contribute to better understanding environmentally-driven MPN clonal evolution, offering a path towards therapeutically preventing their leukemic transformation.
Studying the niche role in AML development and maintenance
AML development and maintenance are complex processes that are only partially explained by the intrinsic accumulation of genetic alterations in hematopoietic stem and progenitor cells. Recent findings have demonstrated the key role of the bone marrow niche in sustaining AML and regulating drug resistance.
The goal of our work is to define and validate novel niche-leukemic crosstalk induced dependencies, through combining single cell “omic” approaches to innovative and powerful high-throughput large scale screening tools applied to physiologically relevant custom made in vitro and in vivo models. We also develop a pipeline to validate the identified therapeutic targets and conduct the necessary preclinical steps towards translating our findings to the clinic, including identification of novel biomarkers of response to treatments and synergy studies. In parallel, we are highly interested in shedding light on the mechanistic underpinnings of the bone marrow-leukemia crosstalk using transcriptomic-, epigenomic- and in vivo microscopy-based approaches.
Deciphering the pathways involved in the leukemic-niche protective partnership will open the road for concomitant “seed” and “soil” targeting strategies, resulting in AML leukemic stem cells eradication and patient’s survival improvement.
ZAFRANI group
Exploring acute life-threatening complications of acute myeloid leukemia
Deciphering the mechanisms of AML-related organ failures (tumor lysis syndrome, leukostasis)
Five to 20% of patients with AML initially present with a high level of circulating leukocytes, exceeding 50-100 x109 /L. These patients are at high risk of acute life-threatening complications related to tissue infiltration by blast cells, leukostasis or tumor lysis syndrome. Our team has demonstrated that endothelial dysfunction is central in acute kidney injury related to tumor lysis syndrome. Moreover, through the expression of different adhesion molecules, interactions between leukemic and endothelial cells seem to play an important role in acute respiratory failure in AML patients. By using in vivo models of AML and intravital microscopy, we aim to explore endothelial dysfunction in different organs that may be injured in hyperleukocytic AML. Deciphering the mechanisms of this bidirectional dialogue between leukemic cells and endothelial cells will reveal possible therapeutic targets to be explored to improve the survival of AML patient.
Studying the impact of sepsis on AML microenvironment, leukemic cells proliferation and disease progression
Patients with AML are at higher risk of early death due to infectious complications that often require Intensive Care Unit admission. Indeed, the risk of sepsis in patients with hematological malignancies has been estimated to be 15 times higher than that in the general population. Among cancer patients, patients with AML who developed sepsis have a much higher mortality rate than other patients. Acute bacterial infections may alter the tissue microenvironment followed by the path of tumorigenesis in the hematopoietic system. Moreover, inflammatory signals generated by the immune response to sepsis may directly impact hematopoietic stem cells. Circulating cytokines appear to reach hematopoietic stem cells in their niche via the systemic circulation, leading to cell-autonomous effects and the survival of AML cells. Pro-inflammatory cytokines critically regulate hematopoiesis, hematopoietic progenitors, and the bone marrow niche. By inducing sepsis in relevant in vitro and in vivo AML and MPN models, we aim to explore essential cellular and molecular signals that may be triggered by sepsis-induced inflammation, in order to understand the interaction between sepsis-induced inflammation and the AML microenvironment that may exert a strong influence on the progression of leukemia.
► Ellegast JM, Alexe G, Hamze A, Lin S, Uckelmann HJ, Rauch PJ, Pimkin M, Ross LS, Dharia NV, Robichaud AL, Conway AS, Khalid D, Perry JA, Wunderlich M, Benajiba L, Pikman Y, Nabet B, Gray NS, Orkin SH, Stegmaier K. Unleashing Cell-Intrinsic Inflammation as a Strategy to Kill AML Blasts. Cancer Discov. 2022 Jul 6;12(7):1760-1781. doi: 10.1158/2159-8290.CD-21-0956. PMID: 35405016; PMCID: PMC9308469.
► Lin KH, Rutter JC, Xie A, Killarney ST, Vaganay C, Benaksas C, Ling F, Sodaro G, Meslin PA, Bassil CF, Fenouille N, Hoj J, Washart R, Ang HX, Cerda-Smith C, Chaintreuil P, Jacquel A, Auberger P, Forget A, Itzykson R, Lu M, Lin J, Pierobon M, Sheng Z, Li X, Chilkoti A, Owzar K, Rizzieri DA, Pardee TS, Benajiba L, Petricoin E, Puissant A, Wood KC. P2RY2-AKT activation is a therapeutically actionable consequence of XPO1 inhibition in acute myeloid leukemia. Nat Cancer. 2022 Jul;3(7):837-851. doi: 10.1038/s43018-022-00394-x. Epub 2022 Jun 6. PMID: 35668193.
► Benajiba L, Kiladjian JJ. The challenge of targets and drug discovery using large-scale screening approaches in onco-hematology. Therapie. 2022 Mar-Apr;77(2):151-155. doi: 10.1016/j.therap.2021.11.007. Epub 2021 Nov 25. PMID: 34895756.
► Fodil S, Arnaud M, Vaganay C, Puissant A, Lengline E, Mooney N, Itzykson R, Zafrani L. Endothelial cells: major players in acute myeloid leukaemia. Blood Rev. 2022 Jul;54:100932. doi: 10.1016/j.blre.2022.100932.
► Arnaud M, Loiselle M, Vaganay C, Pons S, Letavernier E, Demonchy J, Fodil S, Nouacer M, Placier S, Frère P, Arrii E, Lion J, Mooney N, Itzykson R, Djediat C, Puissant A, Zafrani L. Tumor Lysis Syndrome and AKI: Beyond Crystal Mechanisms. J Am Soc Nephrol. 2022 Jun;33(6):1154-1171
► Abdel-Nabey M, Chaba A, Serre J, Lengliné E, Azoulay E, Darmon M, Zafrani L. Tumor lysis syndrome, acute kidney injury and disease-free survival in critically ill patients requiring urgent chemotherapy. Ann Intensive Care. 2022 Feb 15;12(1):15.
► Kapandji N, Azoulay E, Zafrani L. Recent advances in neutropenic enterocolitis: Insights into the role of gut microbiota. Blood Rev. 2022 Jul;54:100944.
► Roux B, Vaganay C, Vargas JD, Alexe G, Benaksas C, Pardieu B, Fenouille N, Ellegast JM, Malolepsza E, Ling F, Sodaro G, Ross L, Pikman Y, Conway AS, Tang Y, Wu T, Anderson DJ, Le Moigne R, Zhou HJ, Luciano F, Hartigan CR, Galinsky I, DeAngelo DJ, Stone RM, Auberger P, Schenone M, Carr SA, Guirouilh-Barbat J, Lopez B, Khaled M, Lage K, Hermine O, Hemann MT, Puissant A, Stegmaier K, Benajiba L. Targeting acute myeloid leukemia dependency on VCP-mediated DNA repair through a selective second-generation small-molecule inhibitor. Sci Transl Med. 2021 Mar 31;13(587):eabg1168. doi: 10.1126/scitranslmed.abg1168. PMID: 33790022; PMCID: PMC8672851.
► Marcault C, Zhao LP, Maslah N, Verger E, Daltro de Oliveira R, Soret-Dulphy J, Cazaux M, Gauthier N, Roux B, Clappier E, Parquet N, Dosquet C, Réa D, Zini JM, Vainchenker W, Raffoux E, Giraudier S, Kiladjian JJ, Cassinat B, Benajiba L. Impact of NFE2 mutations on AML transformation and overall survival in patients with myeloproliferative neoplasms. Blood. 2021 Nov 25;138(21):2142-2148. doi: 10.1182/blood.2020010402. PMID: 33945619.
► Pasquer H, Tostain M, Kaci N, Roux B, Benajiba L. Descriptive and Functional Genomics in Acute Myeloid Leukemia (AML): Paving the Road for a Cure. Cancers (Basel). 2021 Feb 11;13(4):748. doi: 10.3390/cancers13040748. PMID: 33670178; PMCID: PMC7916915.
► Zhao LP, Boy M, Azoulay C, Clappier E, Sébert M, Amable L, Klibi J, Benlagha K, Espéli M, Balabanian K, Preudhomme C, Marceau-Renaut A, Benajiba L, Itzykson R, Mekinian A, Fain O, Toubert A, Fenaux P, Dulphy N, Adès L. Genomic landscape of MDS/CMML associated with systemic inflammatory and autoimmune disease. Leukemia. 2021 Sep;35(9):2720-2724. doi: 10.1038/s41375-021-01152-1. Epub 2021 Feb 23. PMID: 33623140.
► Su A, Ling F, Vaganay C, Sodaro G, Benaksas C, Dal Bello R, Forget A, Pardieu B, Lin KH, Rutter JC, Bassil CF, Fortin G, Pasanisi J, Antony-Debré I, Alexe G, Benoist JF, Pruvost A, Pikman Y, Qi J, Schlageter MH, Micol JB, Roti G, Cluzeau T, Dombret H, Preudhomme C, Fenouille N, Benajiba L, Golan HM, Stegmaier K, Lobry C, Wood KC, Itzykson R, Puissant A. The Folate Cycle Enzyme MTHFR Is a Critical Regulator of Cell Response to MYC-Targeting Therapies. Cancer Discov. 2020 Dec;10(12):1894-1911. doi: 10.1158/2159-8290.CD-19-0970. Epub 2020 Aug 21. PMID: 32826232; PMCID: PMC8044910.
► Pons S, Arrii E, Arnaud M, Loiselle M, Ferry J, Nouacer M, Lion J, Cohen S, Mooney N, Zafrani L. Immunomodulation of endothelial cells induced by macrolide therapy in a model of septic stimulation. Immun Inflamm Dis. 2021 Dec;9(4):1656-1669.
► Lemiale V, Pons S, Mirouse A, Tudesq JJ, Hourmant Y, Mokart D, Pène F, Kouatchet A, Mayaux J, Nyunga M, Bruneel F, Meert AP, Borcoman E, Bisbal M, Legrand M, Benoit D, Azoulay E, Darmon M, Zafrani L. Sepsis and Septic Shock in Patients With Malignancies: A Groupe de Recherche Respiratoire en Réanimation Onco-Hématologique Study. Crit Care Med. 2020 Jun;48(6):822-829
► Benajiba L, Alexe G, Su A, Raffoux E, Soulier J, Hemann MT, Hermine O, Itzykson R, Stegmaier K, Puissant A. Creatine kinase pathway inhibition alters GSK3 and WNT signaling in EVI1-positive AML. Leukemia. 2019 Mar;33(3):800-804. doi: 10.1038/s41375-018-0291-x. Epub 2018 Nov 2. PMID: 30390009; PMCID: PMC6596421.
► Wagner FF, Benajiba L, Campbell AJ, Weïwer M, Sacher JR, Gale JP, Ross L, Puissant A, Alexe G, Conway A, Back M, Pikman Y, Galinsky I, DeAngelo DJ, Stone RM, Kaya T, Shi X, Robers MB, Machleidt T, Wilkinson J, Hermine O, Kung A, Stein AJ, Lakshminarasimhan D, Hemann MT, Scolnick E, Zhang YL, Pan JQ, Stegmaier K, Holson EB. Exploiting an Asp-Glu « switch » in glycogen synthase kinase 3 to design paralog-selective inhibitors for use in acute myeloid leukemia. Sci Transl Med. 2018 Mar 7;10(431):eaam8460. doi: 10.1126/scitranslmed.aam8460. PMID: 29515000; PMCID: PMC6553635.
► Fenouille N, Bassil CF, Ben-Sahra I, Benajiba L, Alexe G, Ramos A, Pikman Y, Conway AS, Burgess MR, Li Q, Luciano F, Auberger P, Galinsky I, DeAngelo DJ, Stone RM, Zhang Y, Perkins AS, Shannon K, Hemann MT, Puissant A, Stegmaier K. The creatine kinase pathway is a metabolic vulnerability in EVI1-positive acute myeloid leukemia. Nat Med. 2017 Mar;23(3):301-313. doi: 10.1038/nm.4283. Epub 2017 Feb 13. PMID: 28191887; PMCID: PMC5540325.
► Benajiba L, Michot JM, Baldini C, Faivre L, Varga A, Balheda R, Gazzah A, Ileana E, Postel-Vinay S, Massard C, de Botton S, Soria JC, Ribrag V. Prognostic factors and outcome of patients with hematological malignancies in phase I trials: the Gustave Roussy scoring system. Anticancer Drugs. 2017 Jun;28(5):540-545. doi: 10.1097/CAD.0000000000000487. PMID: 28225458.
► Neumann T, Benajiba L, Göring S, Stegmaier K, Schmidt B. Evaluation of Improved Glycogen Synthase Kinase-3α Inhibitors in Models of Acute Myeloid Leukemia. J Med Chem. 2015 Nov 25;58(22):8907-19. doi: 10.1021/acs.jmedchem.5b01200. Epub 2015 Nov 5. PMID: 26496242; PMCID: PMC4883112.
Hélène CABANAS
Postdoctoral fellow
Current position: Immuno-Oncology scientist at TreeFrog Therapeutics
Maëlys TOSTAIN
Master 1 student
Current position: MDPhD student at Université Paris Cité
Alexia MONIER
Master 1 student
Current position: PhD student at Université Paris Cité
Juliette BONTOUX
Master 1 student
Current position: Master 2 student at EPHE
Safaa MISKI
Summer intern
Current position: Engineering school student at EPISEN